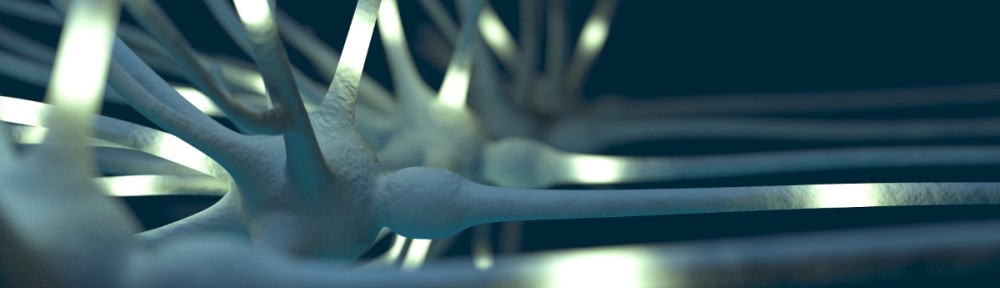
This video is fascinating and frustrating (from a computational modeling point of view) at the same time. Jeff Lichtman shows in this video (above) the complexity of a tiny little fraction of the mouse brain scanned on a nanometer scale using serial electron microscopy. At 16:00 and the following minutes he shows the relation between the nanometer scale, the complexity that can be observed on this level and the large scale view on the brain. Absolutely impressive.
Monthly Archives: May 2014
Interesting Articles (1)
Functional connectivity of the entorhinal-hippocampal space circuit by Sheng-Jia Zhang and colleagues from the Kavli Institute for Systems Neuroscience in Trondheim.
They used light-evoked discharges of grid- and other cells to determine whether these cells have direct hippocampal projections. What I find interesting about this study is that principle cells in MEC need between <5-30ms to fire, depending on the current they receive.
New and Distinct Hippocampal Place Codes Are Generated in a New Environment during Septal Inactivation by Mark Brandon and colleagues from the Center for Functional Connectomics in Seoul.
In this paper they show that septal inactivation leads to reduced theta power and grid cell activity. In a novel environment place fields emerged which suggests (according to their interpretation) that neurons can generate place fields without grid cell activity.
Works related to Novelty Search
Novelty Search has been proposed as alternative to objective driven search in evolutionary runs. It has been found that in some problem domains, Novelty Search performs better than an objective driven search in finding a spot in a large but deceptive solution space. This list links the general idea behind Novelty Search to works from other disciplines which go in a similar direction.
Read VTK files from the Allen Brain Atlas in Blender
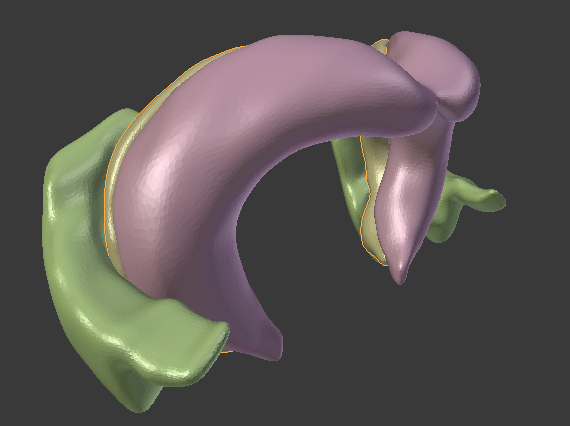
The Allen Brain Atlas provides most of their tracked brain data as 3d-files in the VTK-format. I found it difficult to find a converter for this format that would give me e.g. an obj-file. I came across the Graph Toolbox for Matlab which can read vtk-files and write obj-files. However, their importer did not work for that particular version of the vtk-format. Here is read_Allen_vtk.m, an m-file that I derived from read_vtk.m, which is able to extract all vertices and face-data from the vtk-files. You can then save them using write_obj() and open them in the 3d-tool of your choice. In the upper image you see the hippocampus and entorhinal cortex of the mouse in Blender.