I am proud to announce the start of a new long term project that begins with a paper that came out recently.
Pyka M, Klatt S, Cheng S (2014) Parametric Anatomical Modeling: A method for modeling the anatomical layout of neurons and their projections, Frontiers in Neuroanatomy.
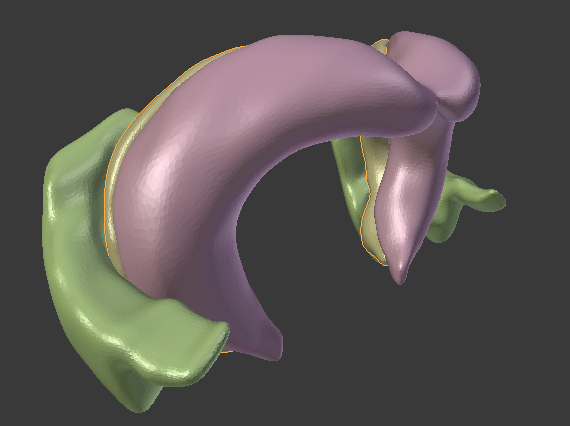
PAM is the foundation for studying generic and specific coding principles of neural networks. The idea is to create artificial neural networks whose connectivity properties and connection lengths result from reconstructed anatomical data. Using a 3d environment to model and describe real neural networks, we can describe complex relationships between layers of neurons. With PAM, we can account for connectivity properties that are influenced by global and local anatomical axes, non-linear relationships between distances of neurons and their connection lengths, and we can incorporate many experimental data, like images from tracer studies, directly into our model.
In the long run, I hope that PAM can help to understand how the structure but also how self-organizing principles (like diverse forms of plasticity) contribute to the topology and thereby also to the function of the network. In particular, we are interested in the functioning of the hippocampus.
The project website is hosted on Bitbucket Github.
Updated on 26.05.2015:
Replaced link to Bitbuckt by link to Github.